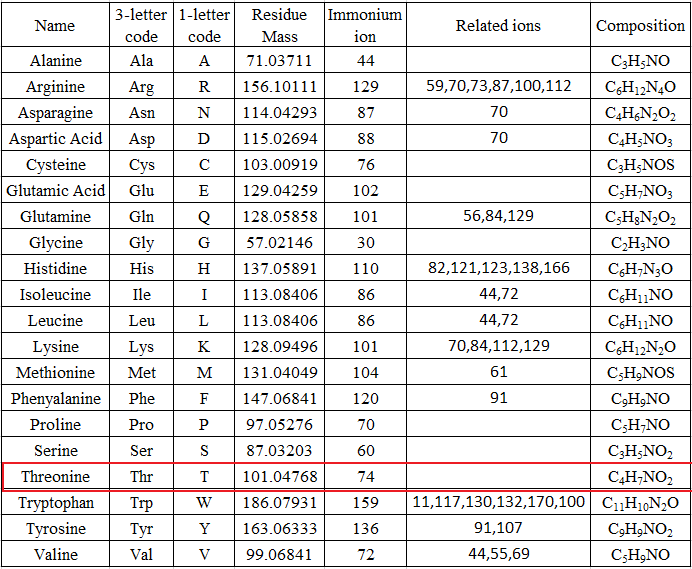
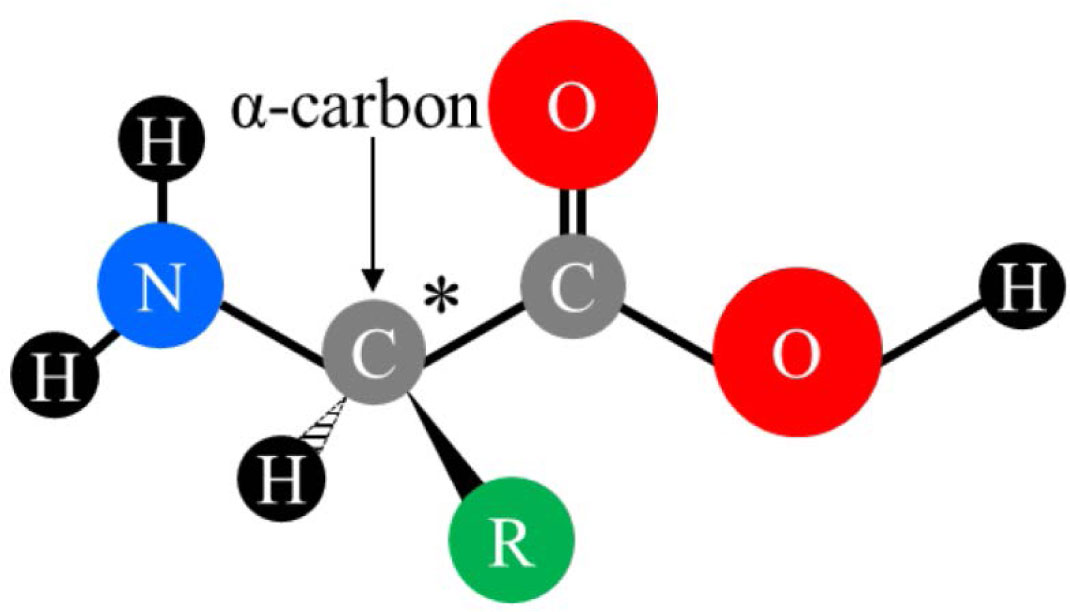
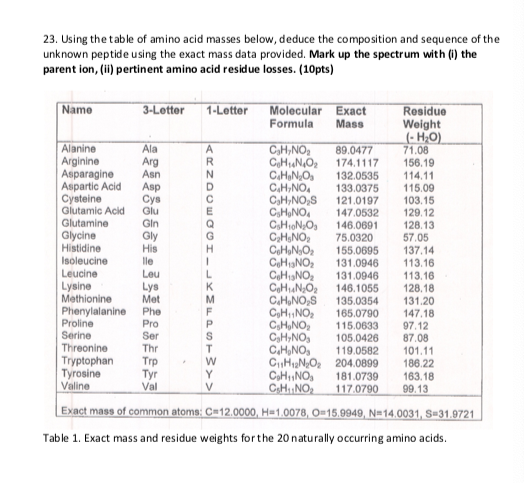
Summary table of the mass, chemical formula, amino acid composition,... | Download Scientific Diagram

Gut amino acid absorption in humans: Concepts and relevance for postprandial metabolism - Clinical Nutrition Open Science

Table 1 from Mass-based classification (MBC) of peptides: Highly accurate precursor ion mass values can be used to directly recognize peptide phosphorylation | Semantic Scholar

Systematic Detection of Amino Acid Substitutions in Proteomes Reveals Mechanistic Basis of Ribosome Errors and Selection for Translation Fidelity - ScienceDirect

Table 2.1 from NBPMF: Novel network-based inference methods for peptide mass fingerprinting | Semantic Scholar

Characterization of chiral amino acids from different milk origins using ultra-performance liquid chromatography coupled to ion-mobility mass spectrometry | Scientific Reports

Expanded Cellular Amino Acid Pools Containing Phosphoserine, Phosphothreonine, and Phosphotyrosine | ACS Chemical Biology
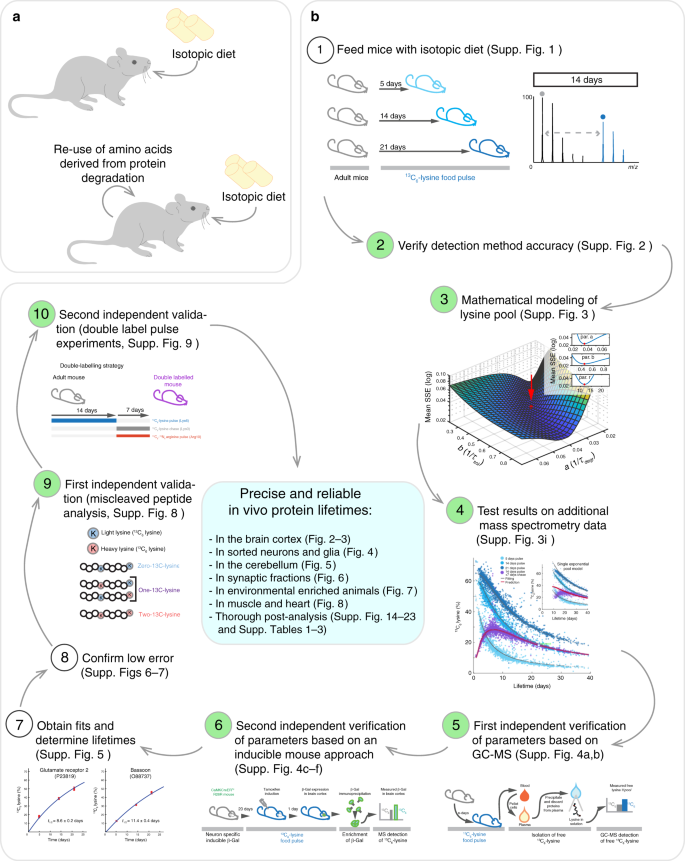
Precisely measured protein lifetimes in the mouse brain reveal differences across tissues and subcellular fractions | Nature Communications
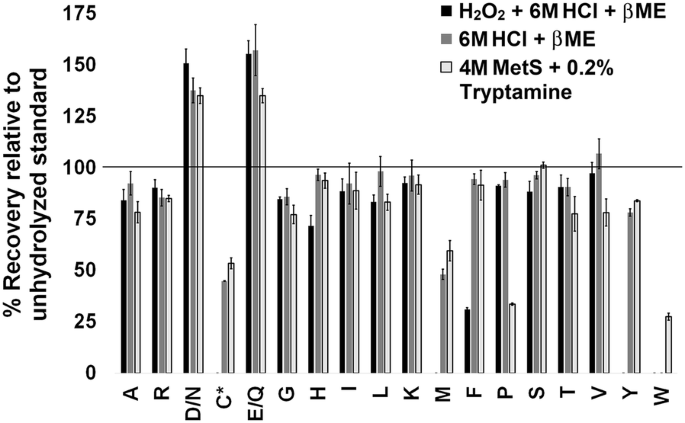
Accurate and efficient amino acid analysis for protein quantification using hydrophilic interaction chromatography coupled tandem mass spectrometry | Plant Methods | Full Text